Re: COVID-19
Posted: Wed May 26, 2021 9:42 am
I wonder if PHE have checked their spreadsheet recently.
The default theme in R/ggplot2 puts minor gridlines uniformly spaced between the major ones, however the scale is transformed. It's not much more of a fiddle to remove them, see, for instance here: https://felixfan.github.io/ggplot2-remo ... nd-margin/.KAJ wrote: ↑Tue May 25, 2021 9:45 pmBecause that's how ggplot2 did it (i.e. I don't know!).
As indicated above, the plot used only 3 ggplot2 functions.Code: Select all
ggplot(subset(UKDF, date > as.Date("2021-04-12")), aes(x = date, y = MVBeds)) + geom_point() + logyAll those functions have many available arguments, most of which I left at the defaults.
- ggplot() specified the data as a subset of the dataframe UKDF, and default x and y variables
- geom_point() added symbols
- I had earlier defined logy <- scale_y_continuous(trans='log10'). Reviewing the help, I could have used the slightly simpler but equivalent logy <- scale_y_log10(). I'll probably change my code to use that.
scale_y_continuous() has an argument `breaks` which I could have specified but left at the default breaks = waiver() which uses "the default breaks computed by the transformation object". If you're really interested () the relevant code may be at https://github.com/r-lib/scales/blob/master/R/breaks-log.R
Cummings is confirming what most of us agreed in March 2020 - that the govt was intending to recklessly and deliberately run Covid hot to diminish future waves.
On 16 March I said:Flatten at a higher level, hoping for huge numbers of non-vulnerable people to catch it, recover and be immune.
The plan is absolutely and almost explicitly stating that the daily infection rate is to be higher than other EU countries. The fundamental idea is the belief they will be low now, then uncontrolled very high later on this year; the UK will be a constant high throughout the year.
You can't defend the UK plan without implying the WHO and nearly all other EU countries are wrong.
Again on 16 March I compared it to running a nuclear reactor hotter:Herd immunity was such a terrible thing to say. Will live on in infamy, in every history text book.
Almost everyone was saying very similar things. There was an interim few days before the official lockdown announcement when the voluntary lockdown was happening - people removing kids from school, staying at home, crowding out the supermarkets. This was the period 10 March to the official announcement on 23 March. On this forum we were remarkably 100% clear during these few days that the govt was deliberately planning to keep running hot.The govt deliberately decided to run the reactor hotter, to churn out more neutrons early on. Ramp up fast, shove in control rods later than they could have done. The hope is more of the fuel is exhausted by the time the control rods degrade later in the year.
Proof?
- on Thursday ill people were only told to isolate for 7 days, when probably infectious for at least 14
- on Thursday the partners and housemates of ill people did not need to isolate, were free to leave the house and take a packed underground train
Their model now tells them it's time to put in some control rods, but not all. They are not lowering the schools control rod yet. They want the reactor to run a little hotter still.
It's a hell of approach. Part of the reason why they are communicating badly is because they don't dare communicate clearly.
Exactly, and *we* were using confirmed deaths, because we knew that case numbers were unreliable. Which really is not rocket science.El Pollo Diablo wrote: ↑Wed May 26, 2021 12:54 pmI'm still absolutely in shock at that two weeks/four weeks thing. I'm fairly far away from being an epidemiologist but I can draw charts with trendlines and logarithmic axes and it was just so clear it was two weeks, I'm amazed that SAGE were so wrong on it.
Nope, but he and his college friends all started working for MAFF just before the 1967 Foot and Mouth outbreak. Which was stressful - the only time anyone (a farmer) pulled a gun (shotgun) on him (and the local copper).
And we saw the same tardy response in the 2000-2001 F&M outbreak. Ignoring lessons learned from animal disease outbreaks 20 years ago seems stupid.jimbob wrote: ↑Wed May 26, 2021 2:44 pmNope, but he and his college friends all started working for MAFF just before the 1967 Foot and Mouth outbreak. Which was stressful - the only time anyone (a farmer) pulled a gun (shotgun) on him (and the local copper).
They also realised that F&M was airborne as opposed to an animal vector (which was the scientific thinking at the time) due to the infected farms following the prevailing wind.
I don't think base_breaks() is in any of the tidyverse packages. You can't add the break at 100 (which lpm seemed to want) without extending the scale that low, and extending the scale automagically adds the break.Bird on a Fire wrote: ↑Wed May 26, 2021 9:17 amYou can use breaks=base_breaks() to get labels at 1, 10, 100 etc, which will put a label close to the current values (which are all close to 100), which might be more similar to other log plots the folk complaining have seen.KAJ wrote: ↑Tue May 25, 2021 9:45 pm
All those functions have many available arguments, most of which I left at the defaults.
scale_y_continuous() has an argument `breaks` which I could have specified but left at the default breaks = waiver() which uses "the default breaks computed by the transformation object".
<snip>
Code: Select all
ggplot(subset(UKDF, date > as.Date("2021-04-12")), aes(x = date, y = MVBeds)) + geom_point() + scale_y_log10(limits = c(100,NA))You're right - turns out I'd defined it elsewhere, with a comment saying I got it from here https://stackoverflow.com/questions/142 ... not-manualKAJ wrote: ↑Wed May 26, 2021 4:27 pmI don't think base_breaks() is in any of the tidyverse packages. You can't add the break at 100 (which lpm seemed to want) without extending the scale that low, and extending the scale automagically adds the break.Bird on a Fire wrote: ↑Wed May 26, 2021 9:17 amYou can use breaks=base_breaks() to get labels at 1, 10, 100 etc, which will put a label close to the current values (which are all close to 100), which might be more similar to other log plots the folk complaining have seen.KAJ wrote: ↑Tue May 25, 2021 9:45 pm
All those functions have many available arguments, most of which I left at the defaults.
scale_y_continuous() has an argument `breaks` which I could have specified but left at the default breaks = waiver() which uses "the default breaks computed by the transformation object".
<snip>MVBeds.pngCode: Select all
ggplot(subset(UKDF, date > as.Date("2021-04-12")), aes(x = date, y = MVBeds)) + geom_point() + scale_y_log10(limits = c(100,NA))
You're right about that as well.
There's been talk of a "race" between the next wave of infections and the vaccination programme. I've been largely down-weighting the recent increase in cases on the basis that the vaccines protect against severe infection better than against infection. A much-above-zero stall in the decline in healthcare numbers, especially MVBeds, suggests that maybe the race is being lost. Here are the numbers from coronavirus.data.gov.uk for the last few weeks - to avoid being sidetracked I've not graphed them.wilsontown wrote: ↑Wed May 26, 2021 10:38 amI suppose at one level it's not all that surprising that hospital numbers would start to level out. The 7 day case average fell to a low of ~2020 on May 2nd and has risen fairly steadily ever since. It is now ~2608. In general the most vulnerable are vaccinated (and more are vaccinated every day) but we're still going to see some people end up in hospital.
Code: Select all
date Admits InPats MVBeds
2021-05-02 119 1282 196
2021-05-03 123 1300 187
2021-05-04 101 1293 181
2021-05-05 117 1236 182
2021-05-06 120 1157 163
2021-05-07 102 1154 159
2021-05-08 108 1135 151
2021-05-09 102 1116 144
2021-05-10 103 1127 149
2021-05-11 105 1104 138
2021-05-12 116 1065 136
2021-05-13 102 998 129
2021-05-14 94 973 124
2021-05-15 90 963 121
2021-05-16 78 938 123
2021-05-17 101 940 125
2021-05-18 124 895 124
2021-05-19 124 915 122
2021-05-20 130 909 123
2021-05-21 117 918 124
2021-05-22 115 882 123
2021-05-23 NA 918 120
2021-05-24 NA 954 121
2021-05-25 NA NA 128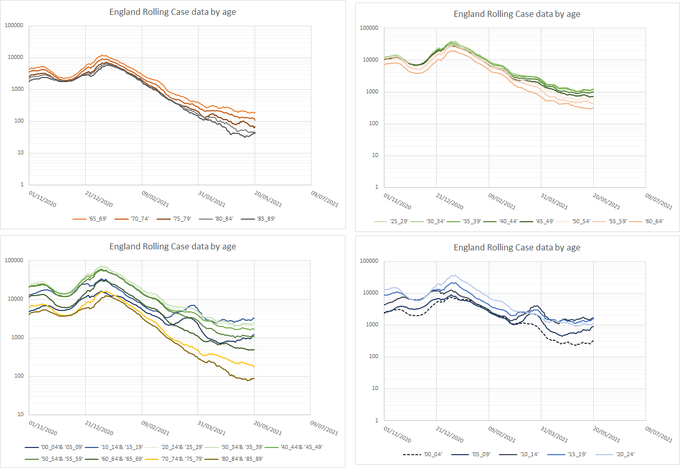
A graph with a meaningless unquantified y-axis made it look like the UK's "flatten the curve in July" strategy would work, until Neil Ferguson quantified it by pointing out that the peak would be many many times higher than the UK's ICU capacity.Bird on a Fire wrote: ↑Wed May 26, 2021 5:30 pmYou're right - turns out I'd defined it elsewhere, with a comment saying I got it from here https://stackoverflow.com/questions/142 ... not-manualKAJ wrote: ↑Wed May 26, 2021 4:27 pmI don't think base_breaks() is in any of the tidyverse packages. You can't add the break at 100 (which lpm seemed to want) without extending the scale that low, and extending the scale automagically adds the break.Bird on a Fire wrote: ↑Wed May 26, 2021 9:17 am
You can use breaks=base_breaks() to get labels at 1, 10, 100 etc, which will put a label close to the current values (which are all close to 100), which might be more similar to other log plots the folk complaining have seen.
<snip>MVBeds.pngCode: Select all
ggplot(subset(UKDF, date > as.Date("2021-04-12")), aes(x = date, y = MVBeds)) + geom_point() + scale_y_log10(limits = c(100,NA))
Quite like your solution too, although personally I do agree that those equidistant minor ticks are a bit confusing.
You're right about that as well.
lpm used to be all in favour of quick-and-dirty early-warning-systems. I'm not sure why this one has caused controversy.
I'll point out that my original graph did have a quantified y-axis, albeit only two labels, and the post included a link to the data.shpalman wrote: ↑Thu May 27, 2021 6:12 amA graph with a meaningless unquantified y-axis made it look like the UK's "flatten the curve in July" strategy would work, until Neil Ferguson quantified it by pointing out that the peak would be many many times higher than the UK's ICU capacity.Bird on a Fire wrote: ↑Wed May 26, 2021 5:30 pmYou're right - turns out I'd defined it elsewhere, with a comment saying I got it from here https://stackoverflow.com/questions/142 ... not-manualKAJ wrote: ↑Wed May 26, 2021 4:27 pm
I don't think base_breaks() is in any of the tidyverse packages. You can't add the break at 100 (which lpm seemed to want) without extending the scale that low, and extending the scale automagically adds the break.MVBeds.pngCode: Select all
ggplot(subset(UKDF, date > as.Date("2021-04-12")), aes(x = date, y = MVBeds)) + geom_point() + scale_y_log10(limits = c(100,NA))
Quite like your solution too, although personally I do agree that those equidistant minor ticks are a bit confusing.
You're right about that as well.
lpm used to be all in favour of quick-and-dirty early-warning-systems. I'm not sure why this one has caused controversy.
So yeah the numbers matter.
My response to lpm's criticism made clear that the plateau was indeed at a value over 100 (emphasis added).Bird on a Fire wrote: ↑Thu May 27, 2021 12:44 pmPlus, the claim was just that the decline had stopped, not that it had stopped at a meaningful value.
You absolutely could verify that from a plot with a "meaningless unquantified y-axis" had one been presented.
But yeah, bit of a fail from ggplot2's default log axis labelling settings there I think. On the first plot I just assumed that major ticks were going in 100s, so you could see that the plateau was at a value over 100, but subsequent plots suggest that's not actually the case.
Somebody tweet Hadley Wickham a link to this thread
KAJ wrote: ↑Tue May 25, 2021 7:24 pmDisagree.lpm wrote: ↑Tue May 25, 2021 5:53 pmThat's a naughty little chart.
Log scale and a short x-axis hide the fact that it's down from 4,000 to 100. While hospital patients are down from 80,000 to 1,000.
It's inevitable that the date will bounce around when a lot levels, while appearing very smooth at high levels. If a wave starts building there won't be anything to see for a couple of weeks due to the lags.
<snip>
The recent data doesn't "bounce around", here are the last 20 reported values in order:My point is that, after a sustained decline, these are remarkably constant - the last 11 are all between 120 and 125Code: Select all
182 163 159 151 144 149 138 136 129 124 121 123 125 124 122 123 124 123 120 121
<snip>
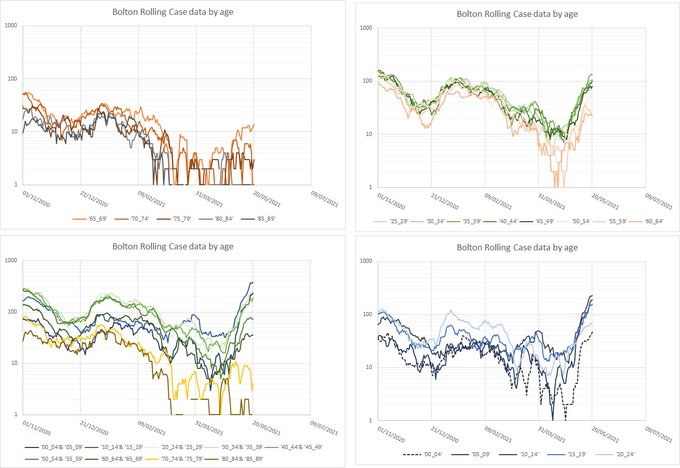
What about Formby?lpm wrote: ↑Thu May 27, 2021 1:56 pmWhat we need is England excluding all towns beginning with B, and the towns beginning with B separately.
Nearly all the country has weeks before it needs to worry about intensive care beds and hospital admissions. A few places will be seeing worrying rises now. The vaccine seems to work just fine against 167-variant hospitalisation so it remains a pretty desperate race to keep ahead.
Just to close (?) the log scales issue, I think Hadley Wickham is aware of the issue! In ggplot2 I found annotation_logticks() which puts minor breaks at the more familiar spacing turning, for example, thisBird on a Fire wrote: ↑Thu May 27, 2021 12:44 pm<snip>
But yeah, bit of a fail from ggplot2's default log axis labelling settings there I think. On the first plot I just assumed that major ticks were going in 100s, so you could see that the plateau was at a value over 100, but subsequent plots suggest that's not actually the case.
Somebody tweet Hadley Wickham a link to this thread
Code: Select all
aDF <- data.frame(x=seq(0,3,0.2), y=10^seq(0,3,0.2))
ggplot(aDF, aes(x = x, y = y)) + geom_point() + scale_y_log10()Code: Select all
ggplot(aDF, aes(x = x, y = y)) + geom_point() + scale_y_log10() + annotation_logticks(sides = "l")